Background
Antibodies are very important in human health. They find and recognize foreign objects (called "antigens") in our bodies and help us fight off infection, especially when we become sick. They work by either preventing viruses or harmful bacteria or other foreign antigens from entering our cells or they actually activate the cells responsible for killing viruses or other pathogens.
Antibodies are very important in human health. They find and recognize foreign objects (called "antigens") in our bodies and help us fight off infection, especially when we become sick. They work by either preventing viruses or harmful bacteria or other foreign antigens from entering our cells or they actually activate the cells responsible for killing viruses or other pathogens.
|
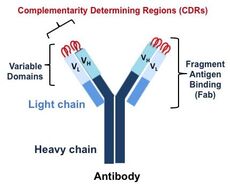
Antibodies have an immunoglobulin form that looks like a Y shaped molecule (Fig. 1). The fragment antigen binding (Fab) region contacts an antigen, such as a bacteria or virus, through its variable domains, which are highly variable. These are the regions that are recombined and mutated to produce a large repertoire of antibodies, which allows the antibodies to recognize so many different foreign antigens. The tips of these molecules have loops, called complementarity determining regions (CDRs), that directly make contact with an antigen. Our lab focuses on the fragment antigen binding (Fab) region, which contacts an antigen, and thus is important for vaccine design studies and for designing antibodies as therapeutics.
|
Figure 1. Antibody Molecule
|
B cells, which produce antibodies, have a receptor (called the B cell receptor or "BCR") on their outer surface, that bind to an antigen. Binding of an antigen results in activation, and a variety of signaling pathways in the cell are turned on, and eventually leading to the proliferation of B cells. B cells also undergo an evolutionary process which is influenced by the antigen that these cells are presented with. This evolutionary process is called antibody affinity maturation and results in B cells produce antibodies with increased affinity for antigen during the course of an immune response.
Vaccines are also capable of activating B cells and producing antibodies. Vaccines are sometimes designed to mimic the antigen (in our case the viral spike) and so they are not infectious but display to the immune system the components necessary for tackling the virus. However, such designs don't always produce the expected antibody response. In order to elicit antibodies via vaccination, we need to better understand how antibodies bind to the viral spikes, how antibodies evolve in response to a vaccine or infection, and how B cells, which produce antibodies, are activated when bound to spikes.
Vaccines are also capable of activating B cells and producing antibodies. Vaccines are sometimes designed to mimic the antigen (in our case the viral spike) and so they are not infectious but display to the immune system the components necessary for tackling the virus. However, such designs don't always produce the expected antibody response. In order to elicit antibodies via vaccination, we need to better understand how antibodies bind to the viral spikes, how antibodies evolve in response to a vaccine or infection, and how B cells, which produce antibodies, are activated when bound to spikes.
Our Areas of Focus
|
Analyzing Interactions between Viral Spikes and Antibodies.
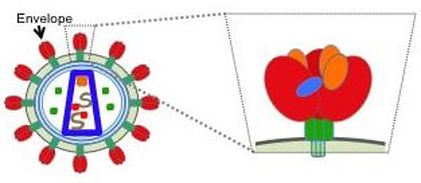
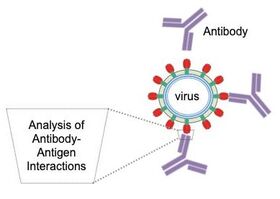
Viral spikes are trimeric assemblies (shown in red and green in the figure above (Fig. 2)) found on the outside of the virus; it makes up the "spike" of the virus, that bind to a molecule outside a host's cell before entering. Much of our focus is on the HIV Envelope (Env) spike (Fig. 2), but we also focus on spikes from other RNA viruses, including the influenza hemagglutinin (HA) and the SARS-CoV-2 spike (S). We study antibodies from infected individuals who have managed to fight off these viruses (or control it in the case of HIV) as well as antibodies from animals that have been immunized with some form of the spike. Specifically, we analyze interactions between these antibodies and the viral spikes (Fig. 3). Analyzing antibodies from donors that could fight off the virus will provide insight on features of the antibodies that are good to elicit by vaccination. Moreover, analyzing antibodies from immunization trials will provide insight on how good the vaccine design strategy was and suggest further improvements. |
Figure 3. Antibodies binding to spikes on viral surface
|
Of note, in the case of rapidly evolving pathogens, such as HIV and flu, a goal of vaccine development is to elicit antibodies that target many viral variants. These antibodies are called broadly neutralizing antibodies (bnAbs). B cells which produce antibodies, undergo an evolutionary process which is influenced by the antigen that these cells are presented with. This evolutionary process is called antibody affinity maturation and results in B cells produce antibodies with increased affinity for antigen during the course of an immune response - this is how bnAbs become potent. Over several rounds of selection, the resultant secreted antibodies produced will have effectively increased affinities for antigen. The result is the production of improved antibodies that effectively recognize infectious agents, and the production of memory B cells. Memory B cells respond more quickly to a second exposure to antigen, or antibody-secreting plasma cells.
So what happens when B cells are presented with a mutating virus? Well, this antibody affinity maturation process repeats, resulting in new antibodies that can recognize the mutated virus. Because HIV mutates so quickly, infected individuals produce populations of both antibodies and viruses, and so we study their co-evolution. In fact, the same affinity maturation process would happen during vaccination! Some vaccines contain portions of the virus, yet are non-infectious, and so an immune response would be launched since the portion of the virus presented would be considered foreign in the host.
So what happens when B cells are presented with a mutating virus? Well, this antibody affinity maturation process repeats, resulting in new antibodies that can recognize the mutated virus. Because HIV mutates so quickly, infected individuals produce populations of both antibodies and viruses, and so we study their co-evolution. In fact, the same affinity maturation process would happen during vaccination! Some vaccines contain portions of the virus, yet are non-infectious, and so an immune response would be launched since the portion of the virus presented would be considered foreign in the host.
Signaling Pathways That Influence Antibody Production.
|
B cells produce antibodies against both foreign antigens, such as viruses, as well as self antigens. In the case of self antigens, there are processes in the body that indicates that the antibodies are binding self and the B cells that produce these antibodies are destroyed. If they are not destroyed, the result is autoimmune disease.
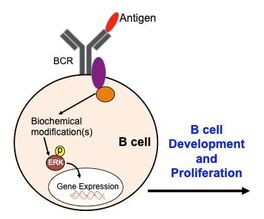
The message that an antigen has bound to receptors on the outer surface of B cells gets transmitted into the cell through signaling pathways involving kinases and other molecules (Fig. 4). Kinases are enzymes that act as molecular on/off switches by transferring a phosphate (negatively charged ion) from one molecule to another. This can turn on other molecules, which then can go on to perform some action, including the expression of certain genes. In the case of B cell activation, B cells proliferate and can become plasma cells which secrete antibodies. Understanding these pathways is important for understanding how B cells function properly to produce antibodies against foreign antigens. This could also provide insights to what makes them fail in eliminating antibodies that bind self antigens. |
Figure 4. Antigen binding to B cell receptor (BCR) activates intracellular proteins via biochemical modifications, such as phosphorylation. This turns on kinases, such as ERK, which can enter the nucleus to turn on transcription factors to turn on genes to lead to cell proliferation.
|
We use approaches from both biochemistry and structural biology to understand the interactions of macromolecules with one another at atomic resolution. We expect that these studies will inform vaccine design strategies and design of antibodies for therapy, and help us better understand the processes that result in antibodies.
We use approaches from both biochemistry and structural biology to understand the interactions of macromolecules with one another at atomic resolution. We expect that these studies will inform vaccine design strategies and design of antibodies for therapy, and help us better understand the processes that result in antibodies.
Techniques Employed
Our lab employs a variety of techniques (the list below indicates some of the main techniques used):
Some examples using our structural approaches are shown in Figures 6-10.
- Sequence Analysis
- DNA recombinant technology
- Protein expression and purification
- Various forms of chromatography (affinity, size exclusion, etc.)
- Electrophoresis
- Bacterial cell growth
- Mammalian tissue culture maintenance
- Pull downs
- Biolayer Interferometry
- Phosphorylation Assays
- X-ray Crystallography
- Negative Stain Electron Microscopy
- Cryo-electron Microscopy
- Computational Modeling
Some examples using our structural approaches are shown in Figures 6-10.
|
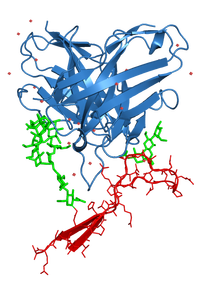
Figure 6. Crystal Structure of an Fab (blue) in complex with a piece of the HIV Envelope (red). Green sticks represent glycans (sugars) found on the HIV envelope surface.
|
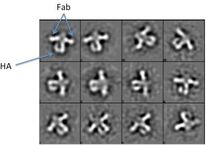
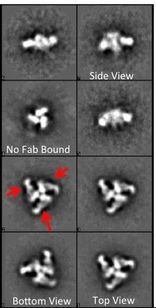
Figure 7. Negative Stain EM 2D Class Averages. Low resolution images can provide us with information on the conformation of HIV Env, as well as where the antibody is binding. Arrows in red are pointing to Fabs on HIV Env.
|
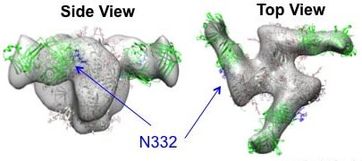
Figure 8. Negative Stain EM 3D Reconstruction. High resolution crystal structures (green and gray ribbon/cartoon) are fit into the low resolution EM map (gray surface). Arrows point to a sugar on HIV Env (at position 332) that is a requirement for the indicated antibody to bind. |
|
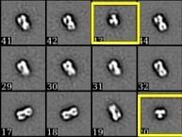
Figure 9. Negative Stain EM 2D Class Averages of HA alone (left) and HA bound to a stem antibody (right). These low resolution images give higher contrast than the raw images that are collected and show how uniform the sample is. They also allow the determination of a 3-dimensional model. Yellow boxes are of "top views" of the HA trimer; the rest are side views.
|
Figure 10. Negative Stain EM 3D Reconstruction. A side view of an HA trimer that was used in mouse immunizations is shown. The next step is to get a model with an antibody bound, either at a similar resolution, or at a higher resolution using cryo-EM.
|
FUNDING
We gratefully acknowledge support from:
- NIH NIAID
- The Pittsburgh Foundation
- Research Corporation for Science Advancement
- amfAR - The Foundation for AIDS Research
- Swarthmore College